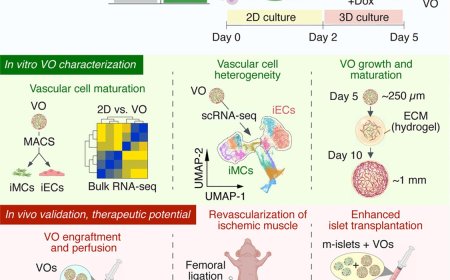
Analyzing genome– environment interactions using medaka model
As a temperate fish species, medaka tolerates fluctuating environments (e.g., seasonal changes and different water salinity levels).
Medaka are highly tolerant to inbreeding; a near-isogenic inbred panel named Medaka Inbred Kiyosu-Karlsruhe panel and several wild-derived inbred strains of medaka have been established.
These panels and strains allow the analysis of phenotype–genotype interactions under different environmental conditions and the modeling of human populations.
Advanced epigenomic approaches, including assay for transposase-accessible chromatin using sequencing, highthroughput chromosome conformation capture (Hi-C), and analysis of histone modifications and DNA methylation, extend genomic approaches in our understanding of gene–environment interactions.
https://www.cell.com/trends/genetics/fulltext/S0168-9525(25)00315-4